Introduction to iSEEpathways
Kevin Rue-Albrecht
University of Oxfordkevin.rue-albrecht@imm.ox.ac.uk
16 October 2024
Source:vignettes/iSEEpathways.Rmd
iSEEpathways.RmdBasics
Install iSEEpathways
R is an open-source statistical environment which can be
easily modified to enhance its functionality via packages. iSEEpathways
is a R package available via the Bioconductor repository for packages.
R can be installed on any operating system from CRAN after which you can install
iSEEpathways
by using the following commands in your R session:
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install("iSEEpathways")
## Check that you have a valid Bioconductor installation
BiocManager::valid()Required knowledge
iSEEpathways is based on many other packages and in particular in those that have implemented the infrastructure needed for dealing with omics data and interactive visualisation. That is, packages like SummarizedExperiment, SingleCellExperiment, iSEE and shiny.
If you are asking yourself the question “Where do I start using Bioconductor?” you might be interested in this blog post.
Asking for help
As package developers, we try to explain clearly how to use our
packages and in which order to use the functions. But R and
Bioconductor have a steep learning curve so it is critical
to learn where to ask for help. The blog post quoted above mentions some
but we would like to highlight the Bioconductor support site
as the main resource for getting help: remember to use the
iSEEpathways tag and check the older
posts. Other alternatives are available such as creating GitHub
issues and tweeting. However, please note that if you want to receive
help you should adhere to the posting
guidelines. It is particularly critical that you provide a small
reproducible example and your session information so package developers
can track down the source of the error.
Citing iSEEpathways
We hope that iSEEpathways will be useful for your research. Please use the following information to cite the package and the overall approach. Thank you!
## Citation info
citation("iSEEpathways")
#> To cite package 'iSEEpathways' in publications use:
#>
#> Rue-Albrecht K, Soneson C (2024). _iSEEpathways: iSEE extension for
#> panels related to pathway analysis_. R package version 1.3.1,
#> <https://github.com/iSEE/iSEEpathways>.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Manual{,
#> title = {iSEEpathways: iSEE extension for panels related to pathway analysis},
#> author = {Kevin Rue-Albrecht and Charlotte Soneson},
#> year = {2024},
#> note = {R package version 1.3.1},
#> url = {https://github.com/iSEE/iSEEpathways},
#> }Quick start to using to iSEEpathways
library("iSEEpathways")
library("fgsea")
library("iSEE")
# Example data ----
set.seed(1)
simulated_data <- simulateExampleData()
pathways_list <- simulated_data[["pathwaysList"]]
features_stat <- simulated_data[["featuresStat"]]
se <- simulated_data[["summarizedexperiment"]]
# fgsea ----
set.seed(42)
fgseaRes <- fgsea(pathways = pathways_list,
stats = features_stat,
minSize = 15,
maxSize = 500)
fgseaRes <- fgseaRes[order(pval), ]
head(fgseaRes)
#> pathway pval padj log2err ES NES size
#> <char> <num> <num> <num> <num> <num> <int>
#> 1: pathway_3193 0.0001211540 0.3204337 0.5384341 -0.2896765 -1.539403 352
#> 2: pathway_2245 0.0001281735 0.3204337 0.5188481 0.3838463 1.789193 119
#> 3: pathway_1942 0.0002489896 0.3688884 0.4984931 0.6384197 2.035004 22
#> 4: pathway_2564 0.0004065213 0.3688884 0.4984931 -0.2960470 -1.536240 266
#> 5: pathway_3993 0.0005506760 0.3688884 0.4772708 -0.4879636 -1.879389 46
#> 6: pathway_4437 0.0005542489 0.3688884 0.4772708 -0.2680266 -1.444987 428
#> leadingEdge
#> <list>
#> 1: feature_....
#> 2: feature_....
#> 3: feature_....
#> 4: feature_....
#> 5: feature_....
#> 6: feature_....
# iSEE ---
se <- embedPathwaysResults(fgseaRes, se, name = "fgsea", class = "fgsea", pathwayType = "simulated",
pathwaysList = pathways_list, featuresStats = features_stat)
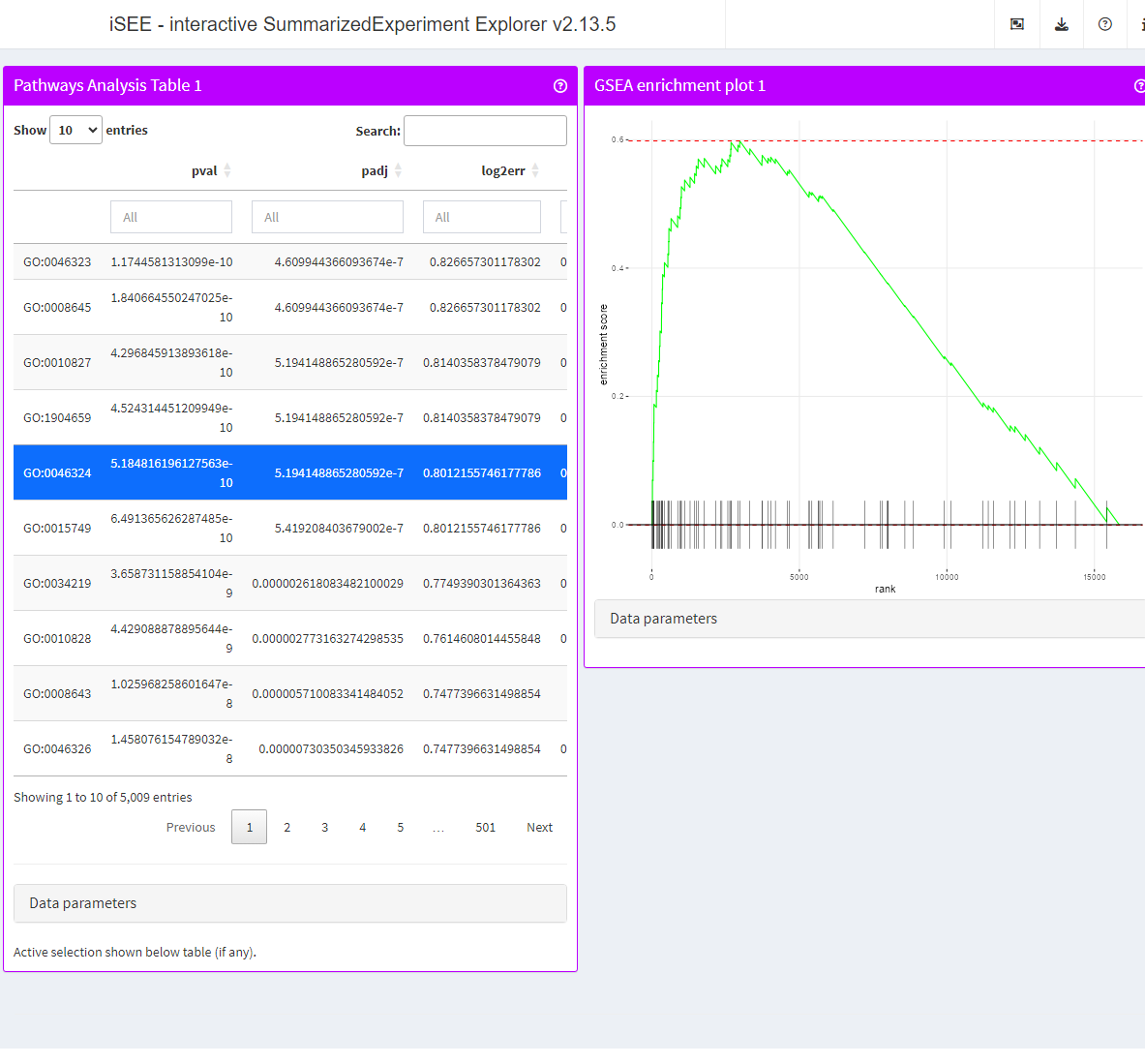
app <- iSEE(se, initial = list(
PathwaysTable(ResultName="fgsea", Selected = "pathway_1350 ", PanelWidth = 6L),
FgseaEnrichmentPlot(ResultName="fgsea", PathwayId = "pathway_1350", PanelWidth = 6L)
))
if (interactive()) {
shiny::runApp(app)
}
Reproducibility
The iSEEpathways package (Rue-Albrecht and Soneson, 2024) was made possible thanks to:
- R (R Core Team, 2024)
- BiocStyle (Oleś, 2024)
- knitr (Xie, 2024)
- RefManageR (McLean, 2017)
- rmarkdown (Allaire, Xie, Dervieux, McPherson, Luraschi, Ushey, Atkins, Wickham, Cheng, Chang, and Iannone, 2024)
- sessioninfo (Wickham, Chang, Flight, Müller, and Hester, 2021)
- testthat (Wickham, 2011)
This package was developed using biocthis.
Code for creating the vignette
## Create the vignette
library("rmarkdown")
system.time(render("iSEEpathways.Rmd", "BiocStyle::html_document"))
## Extract the R code
library("knitr")
knit("iSEEpathways.Rmd", tangle = TRUE)Date the vignette was generated.
#> [1] "2024-10-16 14:48:50 UTC"Wallclock time spent generating the vignette.
#> Time difference of 14.41 secsR session information.
#> ─ Session info ───────────────────────────────────────────────────────────────────────────────────────────────────────
#> setting value
#> version R version 4.4.1 (2024-06-14)
#> os Ubuntu 22.04.5 LTS
#> system x86_64, linux-gnu
#> ui X11
#> language en
#> collate en_US.UTF-8
#> ctype en_US.UTF-8
#> tz UTC
#> date 2024-10-16
#> pandoc 3.4 @ /usr/bin/ (via rmarkdown)
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> abind 1.4-8 2024-09-12 [1] RSPM (R 4.4.0)
#> backports 1.5.0 2024-05-23 [1] RSPM (R 4.4.0)
#> bibtex 0.5.1 2023-01-26 [1] RSPM (R 4.4.0)
#> Biobase * 2.65.1 2024-08-28 [1] Bioconductor 3.20 (R 4.4.1)
#> BiocGenerics * 0.51.3 2024-10-02 [1] Bioconductor 3.20 (R 4.4.1)
#> BiocManager 1.30.25 2024-08-28 [2] CRAN (R 4.4.1)
#> BiocParallel 1.39.0 2024-05-01 [1] Bioconductor 3.20 (R 4.4.0)
#> BiocStyle * 2.33.1 2024-06-12 [1] Bioconductor 3.20 (R 4.4.0)
#> bookdown 0.41 2024-10-16 [1] RSPM (R 4.4.0)
#> bslib 0.8.0 2024-07-29 [2] RSPM (R 4.4.0)
#> cachem 1.1.0 2024-05-16 [2] RSPM (R 4.4.0)
#> circlize 0.4.16 2024-02-20 [1] RSPM (R 4.4.0)
#> cli 3.6.3 2024-06-21 [2] RSPM (R 4.4.0)
#> clue 0.3-65 2023-09-23 [1] RSPM (R 4.4.0)
#> cluster 2.1.6 2023-12-01 [3] CRAN (R 4.4.1)
#> codetools 0.2-20 2024-03-31 [3] CRAN (R 4.4.1)
#> colorspace 2.1-1 2024-07-26 [1] RSPM (R 4.4.0)
#> colourpicker 1.3.0 2023-08-21 [1] RSPM (R 4.4.0)
#> ComplexHeatmap 2.21.1 2024-09-24 [1] Bioconductor 3.20 (R 4.4.1)
#> cowplot 1.1.3 2024-01-22 [1] RSPM (R 4.4.0)
#> crayon 1.5.3 2024-06-20 [2] RSPM (R 4.4.0)
#> data.table 1.16.2 2024-10-10 [1] RSPM (R 4.4.0)
#> DelayedArray 0.31.14 2024-10-03 [1] Bioconductor 3.20 (R 4.4.1)
#> desc 1.4.3 2023-12-10 [2] RSPM (R 4.4.0)
#> digest 0.6.37 2024-08-19 [2] RSPM (R 4.4.0)
#> doParallel 1.0.17 2022-02-07 [1] RSPM (R 4.4.0)
#> dplyr 1.1.4 2023-11-17 [1] RSPM (R 4.4.0)
#> DT 0.33 2024-04-04 [1] RSPM (R 4.4.0)
#> evaluate 1.0.1 2024-10-10 [2] RSPM (R 4.4.0)
#> fansi 1.0.6 2023-12-08 [2] RSPM (R 4.4.0)
#> fastmap 1.2.0 2024-05-15 [2] RSPM (R 4.4.0)
#> fastmatch 1.1-4 2023-08-18 [1] RSPM (R 4.4.0)
#> fgsea * 1.31.6 2024-10-09 [1] Bioconductor 3.20 (R 4.4.1)
#> fontawesome 0.5.2 2023-08-19 [2] RSPM (R 4.4.0)
#> foreach 1.5.2 2022-02-02 [1] RSPM (R 4.4.0)
#> fs 1.6.4 2024-04-25 [2] RSPM (R 4.4.0)
#> generics 0.1.3 2022-07-05 [1] RSPM (R 4.4.0)
#> GenomeInfoDb * 1.41.2 2024-10-02 [1] Bioconductor 3.20 (R 4.4.1)
#> GenomeInfoDbData 1.2.13 2024-10-16 [1] Bioconductor
#> GenomicRanges * 1.57.2 2024-10-09 [1] Bioconductor 3.20 (R 4.4.1)
#> GetoptLong 1.0.5 2020-12-15 [1] RSPM (R 4.4.0)
#> ggplot2 3.5.1 2024-04-23 [1] RSPM (R 4.4.0)
#> ggrepel 0.9.6 2024-09-07 [1] RSPM (R 4.4.0)
#> GlobalOptions 0.1.2 2020-06-10 [1] RSPM (R 4.4.0)
#> glue 1.8.0 2024-09-30 [2] RSPM (R 4.4.0)
#> gtable 0.3.5 2024-04-22 [1] RSPM (R 4.4.0)
#> highr 0.11 2024-05-26 [2] RSPM (R 4.4.0)
#> htmltools 0.5.8.1 2024-04-04 [2] RSPM (R 4.4.0)
#> htmlwidgets 1.6.4 2023-12-06 [2] RSPM (R 4.4.0)
#> httpuv 1.6.15 2024-03-26 [2] RSPM (R 4.4.0)
#> httr 1.4.7 2023-08-15 [1] RSPM (R 4.4.0)
#> igraph 2.0.3 2024-03-13 [1] RSPM (R 4.4.0)
#> IRanges * 2.39.2 2024-07-17 [1] Bioconductor 3.20 (R 4.4.1)
#> iSEE * 2.17.4 2024-09-03 [1] Bioconductor 3.20 (R 4.4.1)
#> iSEEpathways * 1.3.1 2024-10-16 [1] Bioconductor
#> iterators 1.0.14 2022-02-05 [1] RSPM (R 4.4.0)
#> jquerylib 0.1.4 2021-04-26 [2] RSPM (R 4.4.0)
#> jsonlite 1.8.9 2024-09-20 [2] RSPM (R 4.4.0)
#> knitr 1.48 2024-07-07 [2] RSPM (R 4.4.0)
#> later 1.3.2 2023-12-06 [2] RSPM (R 4.4.0)
#> lattice 0.22-6 2024-03-20 [3] CRAN (R 4.4.1)
#> lifecycle 1.0.4 2023-11-07 [2] RSPM (R 4.4.0)
#> listviewer 4.0.0 2023-09-30 [1] RSPM (R 4.4.0)
#> lubridate 1.9.3 2023-09-27 [1] RSPM (R 4.4.0)
#> magrittr 2.0.3 2022-03-30 [2] RSPM (R 4.4.0)
#> Matrix 1.7-0 2024-04-26 [3] CRAN (R 4.4.1)
#> MatrixGenerics * 1.17.0 2024-05-01 [1] Bioconductor 3.20 (R 4.4.0)
#> matrixStats * 1.4.1 2024-09-08 [1] RSPM (R 4.4.0)
#> memoise 2.0.1 2021-11-26 [2] RSPM (R 4.4.0)
#> mgcv 1.9-1 2023-12-21 [3] CRAN (R 4.4.1)
#> mime 0.12 2021-09-28 [2] RSPM (R 4.4.0)
#> miniUI 0.1.1.1 2018-05-18 [2] RSPM (R 4.4.0)
#> munsell 0.5.1 2024-04-01 [1] RSPM (R 4.4.0)
#> nlme 3.1-166 2024-08-14 [2] RSPM (R 4.4.0)
#> pillar 1.9.0 2023-03-22 [2] RSPM (R 4.4.0)
#> pkgconfig 2.0.3 2019-09-22 [2] RSPM (R 4.4.0)
#> pkgdown 2.1.1 2024-09-17 [2] RSPM (R 4.4.0)
#> plyr 1.8.9 2023-10-02 [1] RSPM (R 4.4.0)
#> png 0.1-8 2022-11-29 [1] RSPM (R 4.4.0)
#> promises 1.3.0 2024-04-05 [2] RSPM (R 4.4.0)
#> R6 2.5.1 2021-08-19 [2] RSPM (R 4.4.0)
#> ragg 1.3.3 2024-09-11 [2] RSPM (R 4.4.0)
#> RColorBrewer 1.1-3 2022-04-03 [1] RSPM (R 4.4.0)
#> Rcpp 1.0.13 2024-07-17 [2] RSPM (R 4.4.0)
#> RefManageR * 1.4.0 2022-09-30 [1] RSPM (R 4.4.0)
#> rintrojs 0.3.4 2024-01-11 [1] RSPM (R 4.4.0)
#> rjson 0.2.23 2024-09-16 [1] RSPM (R 4.4.0)
#> rlang 1.1.4 2024-06-04 [2] RSPM (R 4.4.0)
#> rmarkdown 2.28 2024-08-17 [2] RSPM (R 4.4.0)
#> S4Arrays 1.5.11 2024-10-14 [1] Bioconductor 3.20 (R 4.4.1)
#> S4Vectors * 0.43.2 2024-07-17 [1] Bioconductor 3.20 (R 4.4.1)
#> sass 0.4.9 2024-03-15 [2] RSPM (R 4.4.0)
#> scales 1.3.0 2023-11-28 [1] RSPM (R 4.4.0)
#> sessioninfo * 1.2.2 2021-12-06 [2] RSPM (R 4.4.0)
#> shape 1.4.6.1 2024-02-23 [1] RSPM (R 4.4.0)
#> shiny 1.9.1 2024-08-01 [2] RSPM (R 4.4.0)
#> shinyAce 0.4.2 2022-05-06 [1] RSPM (R 4.4.0)
#> shinydashboard 0.7.2 2021-09-30 [1] RSPM (R 4.4.0)
#> shinyjs 2.1.0 2021-12-23 [1] RSPM (R 4.4.0)
#> shinyWidgets 0.8.7 2024-09-23 [1] RSPM (R 4.4.0)
#> SingleCellExperiment * 1.27.2 2024-05-24 [1] Bioconductor 3.20 (R 4.4.0)
#> SparseArray 1.5.44 2024-10-06 [1] Bioconductor 3.20 (R 4.4.1)
#> stringi 1.8.4 2024-05-06 [2] RSPM (R 4.4.0)
#> stringr 1.5.1 2023-11-14 [2] RSPM (R 4.4.0)
#> SummarizedExperiment * 1.35.4 2024-10-09 [1] Bioconductor 3.20 (R 4.4.1)
#> systemfonts 1.1.0 2024-05-15 [2] RSPM (R 4.4.0)
#> textshaping 0.4.0 2024-05-24 [2] RSPM (R 4.4.0)
#> tibble 3.2.1 2023-03-20 [2] RSPM (R 4.4.0)
#> tidyselect 1.2.1 2024-03-11 [1] RSPM (R 4.4.0)
#> timechange 0.3.0 2024-01-18 [1] RSPM (R 4.4.0)
#> UCSC.utils 1.1.0 2024-05-01 [1] Bioconductor 3.20 (R 4.4.0)
#> utf8 1.2.4 2023-10-22 [2] RSPM (R 4.4.0)
#> vctrs 0.6.5 2023-12-01 [2] RSPM (R 4.4.0)
#> vipor 0.4.7 2023-12-18 [1] RSPM (R 4.4.0)
#> viridisLite 0.4.2 2023-05-02 [1] RSPM (R 4.4.0)
#> xfun 0.48 2024-10-03 [2] RSPM (R 4.4.0)
#> xml2 1.3.6 2023-12-04 [2] RSPM (R 4.4.0)
#> xtable 1.8-4 2019-04-21 [2] RSPM (R 4.4.0)
#> XVector 0.45.0 2024-05-01 [1] Bioconductor 3.20 (R 4.4.0)
#> yaml 2.3.10 2024-07-26 [2] RSPM (R 4.4.0)
#> zlibbioc 1.51.1 2024-06-05 [1] Bioconductor 3.20 (R 4.4.0)
#>
#> [1] /__w/_temp/Library
#> [2] /usr/local/lib/R/site-library
#> [3] /usr/local/lib/R/library
#>
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────Bibliography
This vignette was generated using BiocStyle (Oleś, 2024) with knitr (Xie, 2024) and rmarkdown (Allaire, Xie, Dervieux et al., 2024) running behind the scenes.
Citations made with RefManageR (McLean, 2017).
[1] J. Allaire, Y. Xie, C. Dervieux, et al. rmarkdown: Dynamic Documents for R. R package version 2.28. 2024. URL: https://github.com/rstudio/rmarkdown.
[2] M. W. McLean. “RefManageR: Import and Manage BibTeX and BibLaTeX References in R”. In: The Journal of Open Source Software (2017). DOI: 10.21105/joss.00338.
[3] A. Oleś. BiocStyle: Standard styles for vignettes and other Bioconductor documents. R package version 2.33.1. 2024. DOI: 10.18129/B9.bioc.BiocStyle. URL: https://bioconductor.org/packages/BiocStyle.
[4] R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Vienna, Austria, 2024. URL: https://www.R-project.org/.
[5] K. Rue-Albrecht and C. Soneson. iSEEpathways: iSEE extension for panels related to pathway analysis. R package version 1.3.1. 2024. URL: https://github.com/iSEE/iSEEpathways.
[6] H. Wickham. “testthat: Get Started with Testing”. In: The R Journal 3 (2011), pp. 5–10. URL: https://journal.r-project.org/archive/2011-1/RJournal_2011-1_Wickham.pdf.
[7] H. Wickham, W. Chang, R. Flight, et al. sessioninfo: R Session Information. R package version 1.2.2, https://r-lib.github.io/sessioninfo/. 2021. URL: https://github.com/r-lib/sessioninfo#readme.
[8] Y. Xie. knitr: A General-Purpose Package for Dynamic Report Generation in R. R package version 1.48. 2024. URL: https://yihui.org/knitr/.