The goal of iSEEhub is to provide an interface to the Bioconductor ExperimentHub directly within an iSEE web-application.
The main functionality of this package is to define a custom landing page allowing users to browse the Bioconductor ExperimentHub and directly load objects into an iSEE web-application.
Installation instructions
Get the latest stable R release from CRAN. Then install iSEEhub from Bioconductor using the following code:
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install("iSEEhub")And the development version from GitHub with:
BiocManager::install("iSEE/iSEEhub")Example
This is a basic example which shows you how to solve a common problem:
library("iSEEhub")
#> Warning: package 'BiocGenerics' was built under R version 4.2.1
#> Warning: package 'GenomeInfoDb' was built under R version 4.2.1
library(ExperimentHub)
ehub <- ExperimentHub()
app <- iSEEhub(ehub)
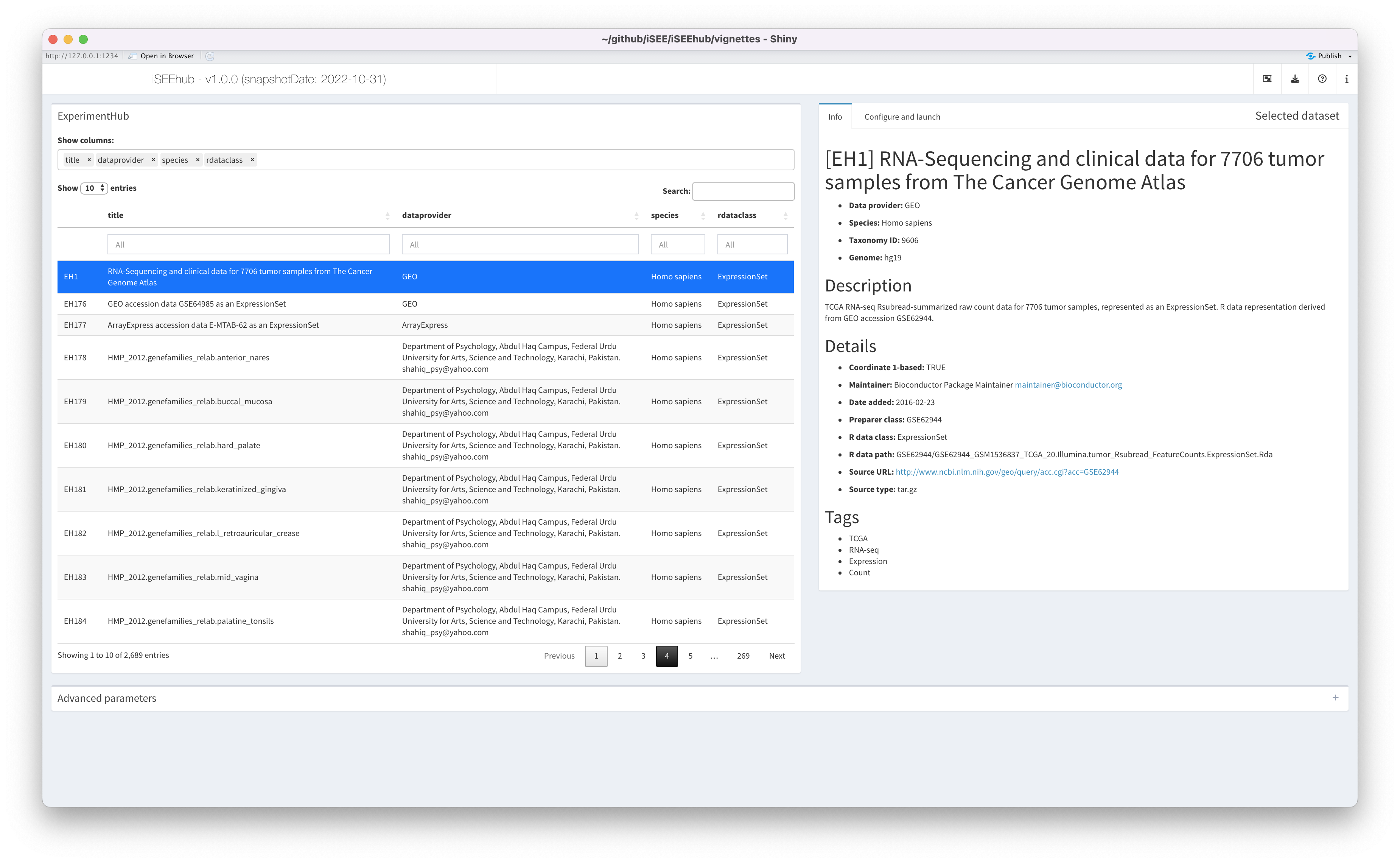
if (interactive()) {
shiny::runApp(app, port = 1234)
}
Citation
Below is the citation output from using citation('iSEEhub') in R. Please run this yourself to check for any updates on how to cite iSEEhub.
print(citation('iSEEhub'), bibtex = TRUE)
#>
#> kevinrue (2022). _Demonstration of a Bioconductor Package_. doi:
#> 10.18129/B9.bioc.iSEEExperimentHub (URL:
#> https://doi.org/10.18129/B9.bioc.iSEEExperimentHub),
#> https://github.com/kevinrue/iSEEExperimentHub/iSEEExperimentHub - R
#> package version 0.99.0, <URL:
#> http://www.bioconductor.org/packages/iSEEExperimentHub>.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Manual{,
#> title = {Demonstration of a Bioconductor Package},
#> author = {{kevinrue}},
#> year = {2022},
#> url = {http://www.bioconductor.org/packages/iSEEExperimentHub},
#> note = {https://github.com/kevinrue/iSEEExperimentHub/iSEEExperimentHub - R package version 0.99.0},
#> doi = {10.18129/B9.bioc.iSEEExperimentHub},
#> }
#>
#> kevinrue (2022). "Demonstration of a Bioconductor Package." _bioRxiv_.
#> doi: 10.1101/TODO (URL: https://doi.org/10.1101/TODO), <URL:
#> https://www.biorxiv.org/content/10.1101/TODO>.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Article{,
#> title = {Demonstration of a Bioconductor Package},
#> author = {{kevinrue}},
#> year = {2022},
#> journal = {bioRxiv},
#> doi = {10.1101/TODO},
#> url = {https://www.biorxiv.org/content/10.1101/TODO},
#> }Please note that the iSEEhub was only made possible thanks to many other R and bioinformatics software authors, which are cited either in the vignettes and/or the paper(s) describing this package.
Code of Conduct
Please note that the iSEEhub project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.
Development tools
- Continuous code testing is possible thanks to GitHub actions through usethis, remotes, and rcmdcheck customized to use Bioconductor’s docker containers and BiocCheck.
- Code coverage assessment is possible thanks to codecov and covr.
- The documentation website is automatically updated thanks to pkgdown.
- The code is styled automatically thanks to styler.
- The documentation is formatted thanks to devtools and roxygen2.
For more details, check the dev directory.
This package was developed using biocthis.
Code of Conduct
Please note that the iSEEhub project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.